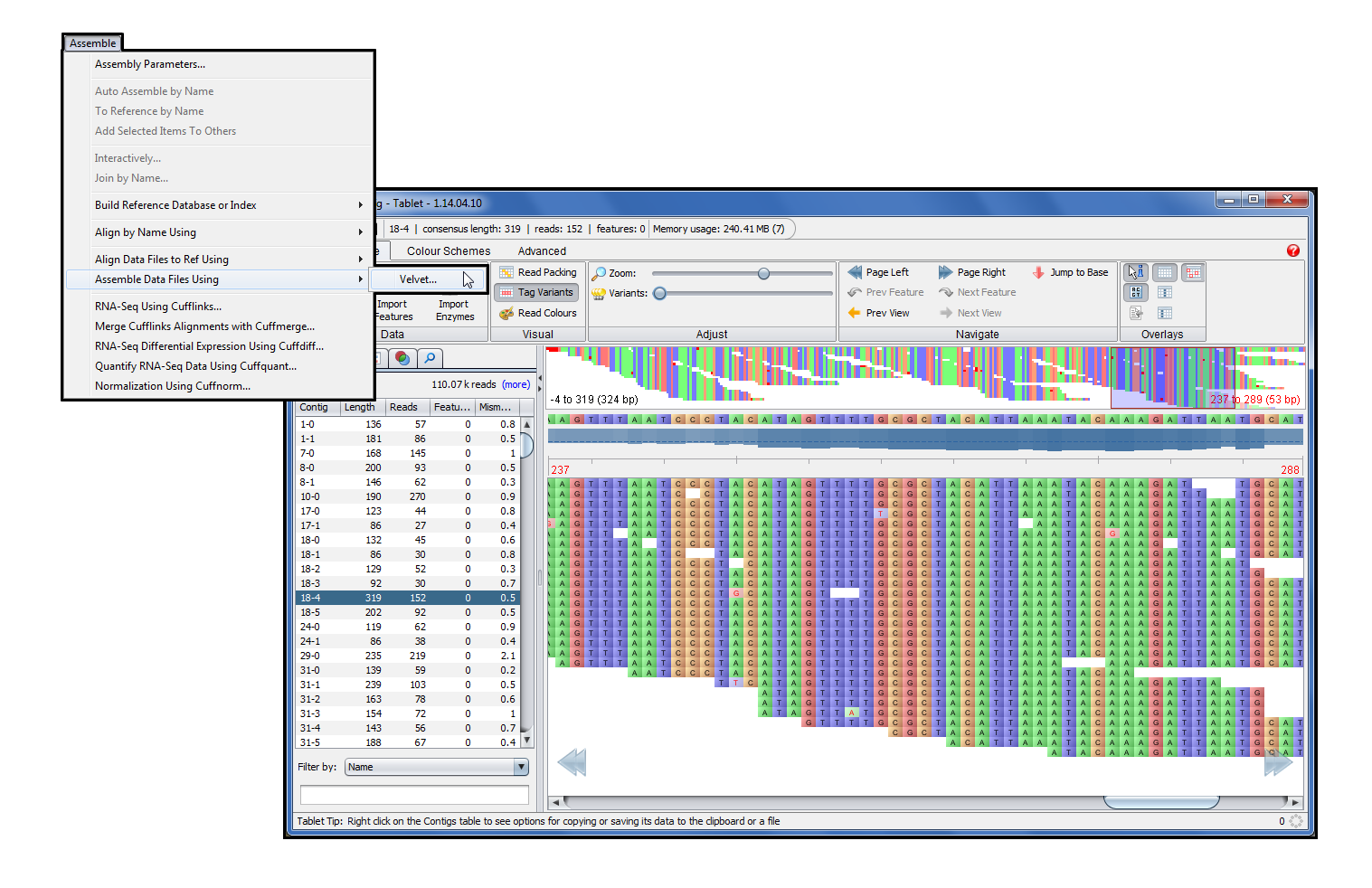
Although possible sinking of certain upper-ocean viruses (e.g. Deep-sea viral sequences were largely uncharacterized, and the majority of 'known' fractions were homologous to Caudovirales. 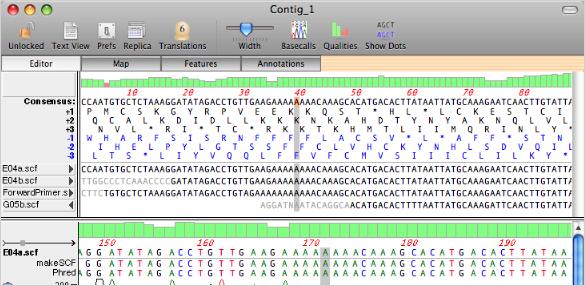
In this study, we performed metagenomics to investigate viral communities from deep sea waters in the South China Sea (SCS) and western Pacific Ocean (WPO). However, despite their importance, viruses in the deep ocean remain largely unknown. The deep ocean, one of the largest biomes on Earth, supports diverse microbial communities, which play important roles in biogeochemical cycles. The approach described should be useful for more detailed investigations of the abundance, dynamics, and sources of DNA in the distinct pools that comprise filterable DNA in aquatic environments. The estimated concentration of DNA that is either F‐DNA, in viruses, or in vesicles was 0.13, 0.14, and 0.08 μg L−1, respectively in the euphotic zone and 0.09, 0.04, and 0.03 μg L−1, respectively in the mesopelagic zone. Application of the fractionation method to seawater samples collected from the oligotrophic North Pacific Ocean followed by analysis of fractions (epifluorescence and electron microscopy, DNase digestion) suggested that the low‐density fractions (1.30–1.35 g mL−1) were dominated by vesicle‐like particles, mid‐density fractions (1.45–1.55 g mL−1) by virus‐like particles, and high‐density fractions (1.60–1.70 g mL−1) by F‐DNA. Spike‐in experiments with a known range of DNA standards (75–20,000 bp) indicated that this method results in high recoveries of F‐DNA (68–86%) with minimal degradation. We investigated whether filtration ( 30 kDa), and fractionation in an equilibrium buoyant density gradient could be used to discriminate the mass contributions of the different pools of filterable DNA in seawater. It is operationally defined as the DNA that passes a membrane filter and thus includes pools of truly dissolved “free” DNA (F‐DNA), virion encapsidated DNA, DNA within membrane vesicles, and possibly other bound forms, each with different sources and lability. Examining microbial processes through the lens of exocellular DNA provides insights into the production of labile dissolved organic matter (i.e., free DNA) at the surface (likely by viral lysis) and processes that influence the fate of sinking, surface-derived organic matter.ĭissolved DNA (D‐DNA) is a ubiquitous component of dissolved organic matter in aquatic systems. Here, we document different microbial sources of free DNA at the surface (0 to 200 m) versus depths of 250 to 1,000 m, suggesting that distinct free DNA production mechanisms may be present throughout the oligotrophic water column. The fate of this free DNA has both ecological consequences as a nutrient (N and P) source and potential evolutionary consequences as a source of genetic transformation. Here, we characterized exocellular free DNA via metagenomics, using a newly developed method that separates free DNA from cells, viruses, and vesicles, and facilitated the independent characterization of each fraction. IMPORTANCE With advances in metagenomic sequencing, the microbial composition of diverse environmental systems has been investigated, providing new perspectives on potential ecological dynamics and dimensions for experimental investigations. These results reveal the composition of free DNA in different regions of the water column (euphotic and mesopelagic zones), with implications for dissolved organic matter cycling and export (by way of sinking particles and/or migratory zooplankton) as a delivery mechanism. Throughout the water column, but especially in the mesopelagic zone, the composition of free DNA sequences was not always reflective of cooccurring microbial communities that inhabit the same sampling depth. A high proportion of mesopelagic zone free DNA sequences appeared to originate from surface waters, including a large amount of DNA contributed by high-light Prochlorococcus ecotypes. Euphotic zone free DNA (75 to 125 m) contained primarily bacterial and viral sequences, with bacteria dominating samples from the mesopelagic zone (500 to 1,000 m). Viral DNA consisted predominantly of myovirus-like and podovirus-like DNA and contained the highest proportion of unannotated sequences. Pelagibacter-like DNA dominated the vesicle fractions for all samples examined over a depth range of 75 to 500 m. Using a method that provides separation of these three fractions, we compared open ocean depth profiles of DNA associated with each fraction. It is composed of DNA-containing vesicles, viruses, and free DNA and is ubiquitous in all aquatic systems, although the sources, sinks, and ecological consequences are largely unknown. Exocellular DNA is operationally defined as the fraction of the total DNA pool that passes through a membrane filter (0.1 μm).
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
